IMGT Repertoire (IG and TR)
Locus representation: human (Homo sapiens) TRB locus on chromosome 7 (7q34)
- Human TRB orphons on chromosome 9 (9p21)
- Polymorphism by insertion/deletion between TRBV4-2 and TRBV7-2
Human (Homo sapiens) TRB locus on chromosome 7 (7q34)
The orientation of the human (Homo sapiens) TRB locus on chromosome 7 (7q34) is forward (FWD).
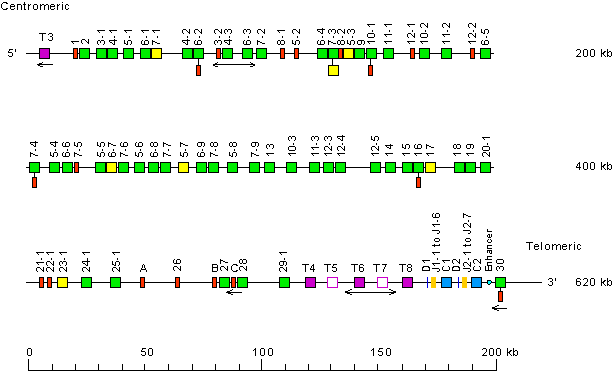
Legend:
Colors are according to IMGT color menu for genes.
The boxes representing the genes are not to scale. Exons are not shown.
A double slash // indicates a gap in genome assembly. Distances in bp (base pair) or in kb (kilobase) associated with a double slash are taken
into account in the length of the lines and included in the numbers displayed at the right end of the lines.
TRBV gene designations according to Rowen et al. (1996). Nomenclature adopted by IMGT.
Single arrows show genes whose polarity is opposite to that of the
D-J-C-CLUSTER.
Double arrows indicate insertion/deletion polymorphisms.
• PRSS58 (serine protease 58, trypsinogen-like TRYX3, TRY1) (5' borne) has been identified 41 kb upstream of TRBV1 (P), the most 5' gene in the locus.
• EPHB6 (EPH receptor B6) (3' borne) has been identified 41 kb downstream of TRBV30 (F), the most 3' gene in the locus.
T3 to T8 indicate trypsinogen or trypsinogen-like genes. T3 (PRSS3P3, TRY3) is at 7.4 kb upstream of TRBV1. T4 to T8 are localized between TRBV29-1 and TRBD1.
Locus representation with only the functional genes: human (Homo sapiens) TRB locus on chromosome 7 (7q34)
Gene localization is according to data from
IMGT/LocusView.
This locus representation with only the functional genes is used with IMGT/GeneFrequency.
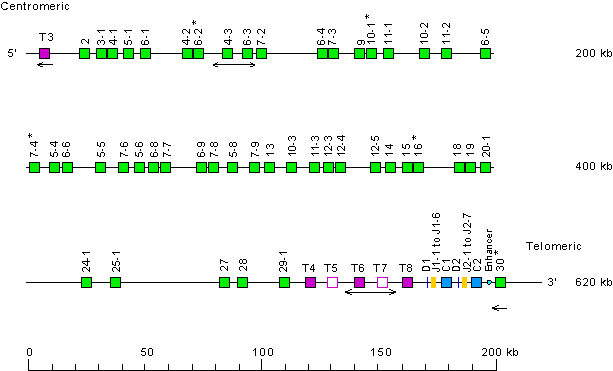
* When needed, indicate the presence of allele(s) with another functionality: see above human (Homo sapiens) TRB locus on chromosome 7 (7q34)
Zoom for TRBD and TRBJ genes

- [1] Lefranc, M.-P. and Lefranc, G., The T cell receptor FactsBook, Academic Press, London, 398 pages (2001).
- [2] Lefranc, M.-P., Nomenclature of the human T cell receptor genes, Current Protocols in Immunology, A.1O.1-A.1O.23 (2000).
- [3] Wei, S. et al., Immunogenetics, 40, 27-36 (1994).
- [4] Posnett, D.N. et al., The Immunologist, 4, 5-8 (1996).
- [5] Rowen, L. et al., Science, 272, 1755-1762 (1996).
- [6] Toyonaga, B. et al., Proc. Natl. Acad. Sci. USA, 82, 8624-8628 (1985).
- [7] Tunnacliffe, A. et al., Proc. Natl. Acad. Sci. USA, 82, 5068-5072 (1985).
- [8] Arden, B. et al., Immunogenetics, 42, 455-500 (1995).
- [9] Gottschalk, L.R. and Leiden, J.M., Mol. Cell. Biol., 10, 5486-5495 (1990).
Links to IMGT/LocusView, IMGT/GENE-DB and genome browsers
- Genome browsers:
- Created:
- 07/10/1996
- Last updated:
- 22/09/2023
- Authors:
- Géraldine Folch
